Our paper on how nucleosomes influence mutation rate leading to a 10bp periodic pattern is out in Cell
We are very excited to share that our work describing the somatic and germline 10bp periodicity in the mutation rate within nucleosome-bound DNA was published in the last 1st of November Cell issue.
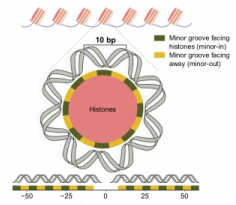
Somatic and Germline Mutation Periodicity Follow the Orientation of the DNA Minor Groove around Nucleosomes
We analyzed the distribution of mutations across the whole genome of more than 3,000 human tumours from The Cancer Genome Atlas (TCGA) and the International Cancer Genome Consortium (ICGC). In several tumor types, we found that mutations accumulate every 10 base pairs (bp) when the DNA is packed around nucleosomes. The phase of the periodicity is different depending on the mutational processes contributing the majority of the mutations in each cancer type. We then explored the whole-genome maps of different types of DNA damage (such as UV light-induced lesions) and repair (such as Nucleotide Excision Repair and Base Excision Repair). The analyzed types of DNA lesions and DNA repair machineries also exhibit a 10 bp periodic pattern, providing a mechanistic explanation for the periodicity observed for the mutations contributed by certain mutational processes. The periodic pattern was also observed in rare gentic variation across human and Arabidopsis populations, and in the genomic sites of divergence of these two species and their closest relatives from their last common ancestor.
The genomes of eukaryotes and certain archeabacteria exhibit a detectable 10bp sequence periodicity consisting of AT, TT, TA, and AA di-nucleotides (WW periodicity). This has long been associated to the presence of nucleosomes. This WW periodicity has been speculated to have arisen through selection of mutations conducive to sequences stabilizing the DNA bending around nucleosomes. Based on our results, we propose that the interplay between damage and repair would contribute to the generation of the WW periodic pattern in the genomes of eukaryotes.
See Núria López-Bigas thread on Tweeter
See also the Cell Preview from Kotler and Segal


