Our paper on reduced mutation rate in exons due to differential mismatch repair published in Nature Genetics
We are happy to announce that our paper on reduced mutation burden in exons caused by differential mismatch repair has been published yesterday, 6th November in Nature Genetics.
Reduced mutation rate in exons due to differential mismatch repair
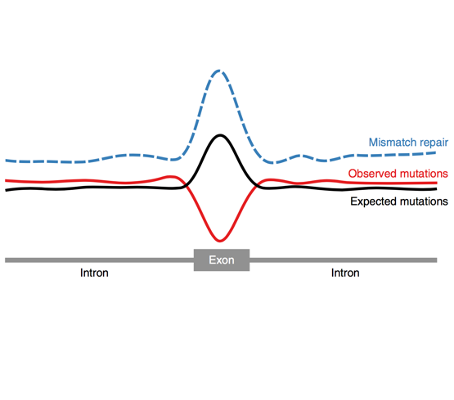
In this study, through the analysis of tumor whole genomes sequenced by The Cancer Genome Atlas (TCGA) consortium, we found that tumors with a preponderance of mismatch-caused mutations show a decrease in exonic mutations with respect to their expected burden. We then demonstrate that this decrease is not due to negative selection. Furthermore, analyzing pediatric brain tumors obtained by The International Biallelic Mismatch Repair Consortium (IBMMRC), we were able to demonstrate that the cause of the observed decrease resides on an increased efficiency of the mismatch repair machinery in exonic regions. Finally, we find evidences that support that differences in chromatin features between exons and introns, more specifically the content of H3K36me3 play a role in this differential mismatch repair function.
We believe these findings have important implications for our understanding of the evolution of eukaryotic genes, as it brings into question the use of rates of intronic substitutions (Ki) as a proxy of neutral evolution. For the same reason, it also has implications for some methods aimed at detecting cancer driver genes that model the background mutation rate of exonic elements by taking as a reference their surrounding areas and, overall, for our understanding of mutational and DNA repair processes.


